InVEST18 | Core Liability Screen
18 high-risk targets with strongest ADR associations. hERG, cardiac ion channels, key GPCRs, transporters. Single-concentration or dose-response.
Read our latest blog: LinkLight™ Protein-Protein Interaction AssaysLearn More
InVEST drug safety screening delivers tiered off-target screening (18, 44, 59, and 77 target panel options) with functional assays and physiological 1mM ATP kinase conditions
Off-target activity is a leading cause of late-stage drug failure. In vitro safety pharmacology profiling evaluates compounds against GPCRs, ion channels, transporters, enzymes, and nuclear receptors clinically linked to adverse drug reactions—enabling discovery teams to design around liabilities before costly in vivo studies.
Reaction Biology’s InVEST platform provides tiered panels from hit confirmation (InVEST18) through extensive profiling (InVEST77). Functional assays distinguish agonist from antagonist activity. Kinases profiled at 1mM ATP reduce false positives. 10 business day turnaround from Malvern, PA.
18 high-risk targets with strongest ADR associations. hERG, cardiac ion channels, key GPCRs, transporters. Single-concentration or dose-response.
Industry-standard 44-target panel (Bowes et al., 2012). Comprehensive coverage: 24 GPCRs, 7 ion channels, 8 enzymes, 3 transporters, 2 nuclear receptors. ICH S7A/B aligned for early hazard identification. Single-concentration or dose-response.
Extended coverage for programs requiring broader safety characterization. All kinases profiled at physiological 1mM ATP to minimize false positives from competitive inhibitors. Supports selectivity-focused SAR studies during structure refinement. Single-concentration or dose-response.
Comprehensive 77-target panel aligned with modern industry standards (Brennan et al., 2024). Expanded kinase coverage at 1mM ATP, additional transporters, and GABA(A) functional assessment. Single-concentration or full dose-response IC50/EC50 available for all targets.
Highly sensitive luminescent detection across 14 CYP isoforms. Reversible inhibition screening, time-dependent inhibition (TDI) assessment, and CYP induction available. Recombinant CYP Supersomes™ ensure consistent enzyme activity.
Comprehensive coverage of 18 phosphodiesterase isoforms for selectivity profiling and off-target liability assessment. Fluorescence-based catalytic assays with Envision plate reader detection. Identifies PDE-related cardiovascular, CNS, and inflammatory liabilities early.
All InVEST panels deliver data in 10 business days, whether screening 18 targets or 77.
One unified InVEST platform (18/44/59/77) enables simple panel selection aligned to your development stage.
Cell-based functional assays for select targets provide mechanism data on agonist/antagonist responses, not just binding. FLIPR calcium flux, manual patch electrophysiology, and reporter gene readouts available.
All assays available in single-concentration format (10 µM or client-requested) or full 8-point dose-response for IC50/EC50 determination.
All kinase targets profiled at physiological 1mM ATP. Reduces false positives from ATP-competitive inhibitors that lack activity at intracellular ATP levels.
All InVEST testing performed at our Malvern, PA facility with our team of safety and toxicology experts.
Pre-planned panel runs enable predictable compound submission dates and data delivery timelines.
| Target Family | Target Name | HGNC Reference | Assay Type | Assay Format | Species | InVEST18 | InVEST44 | InVEST59 | InVEST77 | PDE | CYP |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Other Enzyme | Acetylcholinesterase | ACHE | Biochemical | Enzymatic Activity | Human | ✓ | ✓ | ✓ | |||
| GPCR | Adenosine A2A | ADORA2A | Biochemical | Radioligand Filter Binding | Human | ✓ | ✓ | ✓ | |||
| GPCR | Adrenergic a1A | ADRA1A | Biochemical | Radioligand Filter Binding | Human | ✓ | ✓ | ✓ | |||
| GPCR | Adrenergic a2A | ADRA2A | Biochemical | Radioligand Filter Binding | Human | ✓ | ✓ | ✓ | |||
| GPCR | Adrenergic b1 | ADRB1 | Biochemical | Radioligand Filter Binding | Human | ✓ | ✓ | ✓ | |||
| GPCR | Adrenergic b2 | ADRB2 | Biochemical | Radioligand Filter Binding | Human | ✓ | ✓ | ✓ | |||
| Nuclear Receptor | Androgen (AR) | AR | Cell-based / Functional | Cell Reporter | Human | ✓ | ✓ | ✓ | |||
| Kinase | ATM | ATM | Biochemical | HTRF | Human | ✓ | ✓ | ||||
| Kinase | Aurora A | AURKA | Biochemical | HotSpot | Human | ✓ | ✓ | ||||
| Kinase | Aurora B | AURKB | Biochemical | HotSpot | Human | ✓ | ✓ | ||||
| GPCR | Vasopressin V1A | AVPR1A | Cell-based / Functional | Cell Reporter | Human | ✓ | ✓ | ✓ | |||
| Bromodomain | BRD4 | BRD4 | Biochemical | AlphaScreen Binding | Human | ✓ | ✓ | ||||
| Ion Channel | Cav1.2 | CACNA1C | Cell-based / Functional | Patch Clamp | Human | ✓ | ✓ | ✓ | |||
| GPCR | Cholecystokinin CCK2 | CCKBR | Biochemical | Radioligand Filter Binding | Human | ✓ | ✓ | ||||
| GPCR | Cholecystokinin CCK1 | CCL28 | Biochemical | Radioligand Filter Binding | Human | ✓ | ✓ | ✓ | |||
| GPCR | Muscarinic M1 | CHRM1 | Biochemical | Radioligand Filter Binding | Human | ✓ | ✓ | ✓ | |||
| GPCR | Muscarinic M2 | CHRM2 | Biochemical | Radioligand Filter Binding | Human | ✓ | ✓ | ✓ | |||
| GPCR | Muscarinic M3 | CHRM3 | Biochemical | Radioligand Filter Binding | Human | ✓ | ✓ | ✓ | |||
| Ion Channel | nAChR (a1) | CHRNA1 | Cell-based / Functional | FLIPR/Ca assay | Human | ✓ | ✓ | ||||
| Ion Channel | nAChR (a4/b2) | CHRNA4/CHRNB2 | Cell-based / Functional | FLIPR/Ca Assay | Human | ✓ | |||||
| GPCR | Cannabinoid CB1 | CNR1 | Biochemical | Radioligand Filter Binding | Human | ✓ | ✓ | ✓ | |||
| GPCR | Cannabinoid CB2 | CNR2 | Biochemical | Radioligand Filter Binding | Human | ✓ | ✓ | ✓ | |||
| Protease | Cathepsin D | CTSD | Biochemical | Enzymatic Activity | Human | ✓ | ✓ | ||||
| Cytochrome P450 | CYP19A | CYP19A | Biochemical | Enzymatic Activity | Human | ✓ | |||||
| Cytochrome P450 | CYP1A2 | CYP1A2 | Biochemical | Enzymatic Activity | Human | ✓ | |||||
| Cytochrome P450 | CYP2A6 | CYP2A6 | Biochemical | Enzymatic Activity | Human | ✓ | |||||
| Cytochrome P450 | CYP2B6 | CYP2B6 | Biochemical | Enzymatic Activity | Human | ✓ | |||||
| Cytochrome P450 | CYP2C19 | CYP2C19 | Biochemical | Enzymatic Activity | Human | ✓ | |||||
| Cytochrome P450 | CYP2C8 | CYP2C8 | Biochemical | Enzymatic Activity | Human | ✓ | |||||
| Cytochrome P450 | CYP2C9 | CYP2C9 | Biochemical | Enzymatic Activity | Human | ✓ | |||||
| Cytochrome P450 | CYP2D6 | CYP2D6 | Biochemical | Enzymatic Activity | Human | ✓ | |||||
| Cytochrome P450 | CYP2E1 | CYP2E1 | Biochemical | Enzymatic Activity | Human | ✓ | |||||
| Cytochrome P450 | CYP2J2 | CYP2J2 | Biochemical | Enzymatic Activity | Human | ✓ | |||||
| Cytochrome P450 | CYP3A4 | CYP3A4 | Biochemical | Enzymatic Activity | Human | ✓ | |||||
| Cytochrome P450 | CYP3A5 | CYP3A5 | Biochemical | Enzymatic Activity | Human | ✓ | |||||
| Cytochrome P450 | CYP4A11 | CYP4A11 | Biochemical | Enzymatic Activity | Human | ✓ | |||||
| Cytochrome P450 | CYP4F3B | CYP4F3B | Biochemical | Enzymatic Activity | Human | ✓ | |||||
| GPCR | Dopamine D1 | DRD1 | Biochemical | Radioligand Filter Binding | Human | ✓ | ✓ | ✓ | |||
| GPCR | Dopamine D2L | DRD2 | Biochemical | Radioligand Filter Binding | Human | ✓ | ✓ | ||||
| GPCR | Dopamine D2S | DRD2 | Biochemical | Radioligand Filter Binding | Human | ✓ | |||||
| GPCR | Dopamine D3 | DRD3 | Biochemical | Radioligand Filter Binding | Human | ✓ | ✓ | ||||
| GPCR | Endothelin ETA | EDNRA | Cell-based / Functional | FLIPR/Ca Assay | Human | ✓ | ✓ | ✓ | |||
| Kinase | EGFR | EGFR | Biochemical | HotSpot | Human | ✓ | ✓ | ||||
| Nuclear Receptor | Estrogen-a (ERalpha) | ESR1 | Biochemical | Fluorescence Polarization | Human | ✓ | ✓ | ||||
| Kinase | FGFR1 | FGFR1 | Biochemical | HotSpot | Human | ✓ | ✓ | ||||
| Kinase | FMS (VEGFR1) | FLT1 | Biochemical | HotSpot | Human | ✓ | ✓ | ||||
| Ion Channel | GABAA(A) a1b2g2L | GABRA1/GABRB2/GABRG2 | Cell-based / Functional | FLIPR/Ca Assay | Human | ✓ | |||||
| Ion Channel | GABAA (Central BDZ) | GABRAx | Biochemical | Radioligand Filter Binding | Rat | ✓ | ✓ | ||||
| GPCR | GPBA | GPBAR1 | Cell-based / Functional | Cell Reporter | Human | ✓ | ✓ | ||||
| Kinase | GSK3B | GSK3B | Biochemical | HotSpot | Human | ✓ | ✓ | ||||
| GPCR | Histamine H1 | HRH1 | Biochemical | Radioligand Filter Binding | Human | ✓ | ✓ | ✓ | |||
| GPCR | Histamine H2 | HRH2 | Biochemical | Radioligand Binding | Human | ✓ | ✓ | ✓ | |||
| GPCR | Serotonin 5-HT1A | HTR1A | Biochemical | Radioligand Filter Binding | Human | ✓ | ✓ | ✓ | |||
| GPCR | Serotonin 5-HT1B | HTR1B | Biochemical | Radioligand Filter Binding | Human | ✓ | ✓ | ✓ | |||
| GPCR | Serotonin 5-HT2A | HTR2A | Biochemical | Radioligand Filter Binding | Human | ✓ | ✓ | ✓ | |||
| GPCR | Serotonin 5-HT2B | HTR2B | Biochemical | Radioligand Filter Binding | Human | ✓ | ✓ | ✓ | |||
| Ion Channel | Serotonin 5-HT3 | HTR3A | Biochemical | Radioligand Filter Binding | Human | ✓ | ✓ | ✓ | |||
| GPCR | Serotonin 5-HT6 | HTR6 | Biochemical | Radioligand Filter Binding | Human | ✓ | ✓ | ||||
| Kinase | IR | INSR | Biochemical | HotSpot | Human | ✓ | ✓ | ||||
| Ion Channel | hERG (Kv11.1) | KCNH2 | Biochemical | Fluorescence Polarization | Human | ✓ | ✓ | ✓ | |||
| Ion Channel | KV7.1 | KCNQ1 | Cell-based / Functional | Patch Clamp | Human | ✓ | ✓ | ||||
| Kinase | KDR (VEGFR2) | KDR | Biochemical | HotSpot | Human | ✓ | ✓ | ||||
| Kinase | c-KIT | KIT | Biochemical | HotSpot | Human | ✓ | ✓ | ||||
| Kinase | LCK TK | LCK | Biochemical | ADP-Glo | Human | ✓ | ✓ | ✓ | |||
| Protease | Paracaspase | MALT1 | Biochemical | Enzymatic Activity | Human | ✓ | ✓ | ||||
| MAO | MAO-A | MAOA | Biochemical | Enzymatic Activity | Human | ✓ | ✓ | ✓ | |||
| MAO | MAO-B | MAOB | Biochemical | Enzymatic Activity | Human | ✓ | ✓ | ✓ | |||
| Kinase | MKK7 | MAP2K7 | Biochemical | HotSpot | Human | ✓ | ✓ | ||||
| GPCR | Mas-related GPCR | MRGX2 | Cell-based / Functional | FLIPR/Ca assay | Human | ✓ | ✓ | ||||
| Ion Channel | NMDA | NMDAR1 | Biochemical | Radioligand Filter Binding | Rat | ✓ | ✓ | ✓ | |||
| Nuclear Receptor | Glucocorticoid (GR) | NR3C1 | Biochemical | Fluorescence Polarization | Human | ✓ | ✓ | ✓ | |||
| Kinase | TRKA | NTRK1 | Biochemical | HotSpot | Human | ✓ | ✓ | ||||
| GPCR | Opioid (d) | OPRD1 | Biochemical | Radioligand Filter Binding | Human | ✓ | ✓ | ✓ | |||
| GPCR | Opioid (k) | OPRK1 | Biochemical | Radioligand Filter Binding | Human | ✓ | ✓ | ✓ | |||
| GPCR | Opioid (m) | OPRM1 | Biochemical | Radioligand Filter Binding | Human | ✓ | ✓ | ✓ | |||
| Phosphodiesterase | PDE10A | PDE10A | Biochemical | Enzymatic Activity | Human | ✓ | |||||
| Phosphodiesterase | PDE1A | PDE1A | Biochemical | Enzymatic Activity | Human | ✓ | |||||
| Phosphodiesterase | PDE1B | PDE1B | Biochemical | Enzymatic Activity | Human | ✓ | |||||
| Phosphodiesterase | PDE1C | PDE1C | Biochemical | Enzymatic Activity | Human | ✓ | |||||
| Phosphodiesterase | PDE2A | PDE2A | Biochemical | Enzymatic Activity | Human | ✓ | |||||
| Phosphodiesterase | PDE3A | PDE3A | Biochemical | Enzymatic Activity | Human | ✓ | ✓ | ✓ | ✓ | ||
| Phosphodiesterase | PDE3B | PDE3B | Biochemical | Enzymatic Activity | Human | ✓ | |||||
| Phosphodiesterase | PDE4A | PDE4A | Biochemical | Enzymatic Activity | Human | ✓ | |||||
| Phosphodiesterase | PDE4B | PDE4B | Biochemical | Enzymatic Activity | Human | ✓ | |||||
| Phosphodiesterase | PDE4C | PDE4C | Biochemical | Enzymatic Activity | Human | ✓ | |||||
| Phosphodiesterase | PDE4D | PDE4D | Biochemical | Enzymatic Activity | Human | ✓ | |||||
| Phosphodiesterase | PDE4D2 | PDE4D2 | Biochemical | Enzymatic Activity | Human | ✓ | ✓ | ✓ | ✓ | ||
| Phosphodiesterase | PDE4D2 | PDE4D2 | Biochemical | Enzymatic Activity | Human | ✓ | |||||
| Phosphodiesterase | PDE5A | PDE5A | Biochemical | Enzymatic Activity | Human | ✓ | |||||
| Phosphodiesterase | PDE7A | PDE7A | Biochemical | Enzymatic Activity | Human | ✓ | |||||
| Phosphodiesterase | PDE7B | PDE7B | Biochemical | Enzymatic Activity | Human | ✓ | |||||
| Phosphodiesterase | PDE8A | PDE8A | Biochemical | Enzymatic Activity | Human | ✓ | |||||
| Phosphodiesterase | PDE9A | PDE9A | Biochemical | Enzymatic Activity | Human | ✓ | |||||
| Kinase | PDK1 | PDHK1 | Biochemical | HotSpot | Human | ✓ | ✓ | ||||
| Nuclear Receptor | Progesterone (PR) | PGR | Biochemical | Fluorescence Polarization | Human | ✓ | ✓ | ||||
| Kinase | PI3Kg (p110g) | PIK3CG | Biochemical | ADP-Glo | Human | ✓ | ✓ | ||||
| COX | COX1 | PTGS1 | Biochemical | Enzymatic Activity | Ovine | ✓ | ✓ | ✓ | |||
| COX | COX2 | PTGS2 | Biochemical | Enzymatic Activity | Human | ✓ | ✓ | ✓ | |||
| Kinase | PTK2 | PTK2 | Biochemical | HotSpot | Human | ✓ | ✓ | ||||
| Nuclear Receptor | Retinoic Acid (RARalpha) | RARA | Cell-based / Functional | Cell Reporter | Human | ✓ | ✓ | ||||
| Kinase | ROCK1 | ROCK1 | Biochemical | HotSpot | Human | ✓ | ✓ | ||||
| Kinase | ROCK2 | ROCK2 | Biochemical | HotSpot | Human | ✓ | ✓ | ||||
| Ion Channel | Nav1.5 | SCN5A | Cell-based / Functional | Patch Clamp | Human | ✓ | ✓ | ✓ | |||
| Transporter | GAT1 | SLC6A1 | Biochemical | Radioligand Filter Binding | Human | ✓ | ✓ | ||||
| Transporter | Norepinephrine NET | SLC6A2 | Biochemical | Radioligand Filter Binding | Human | ✓ | ✓ | ✓ | |||
| Transporter | Dopamine DAT | SLC6A3 | Biochemical | Radioligand Filter Binding | Human | ✓ | ✓ | ✓ | |||
| Transporter | Serotonin SERT | SLC6A4 | Biochemical | Radioligand Filter Binding | Human | ✓ | ✓ | ✓ | |||
| Kinase | SRC | SRC | Biochemical | HotSpot | Human | ✓ | ✓ | ||||
| Kinase | STK35 | STK35 | Biochemical | HotSpot | Human | ✓ | ✓ | ||||
| GPCR | NK3 receptor | TACR3 | Biochemical | Radioligand Filter Binding | Human | ✓ | ✓ |
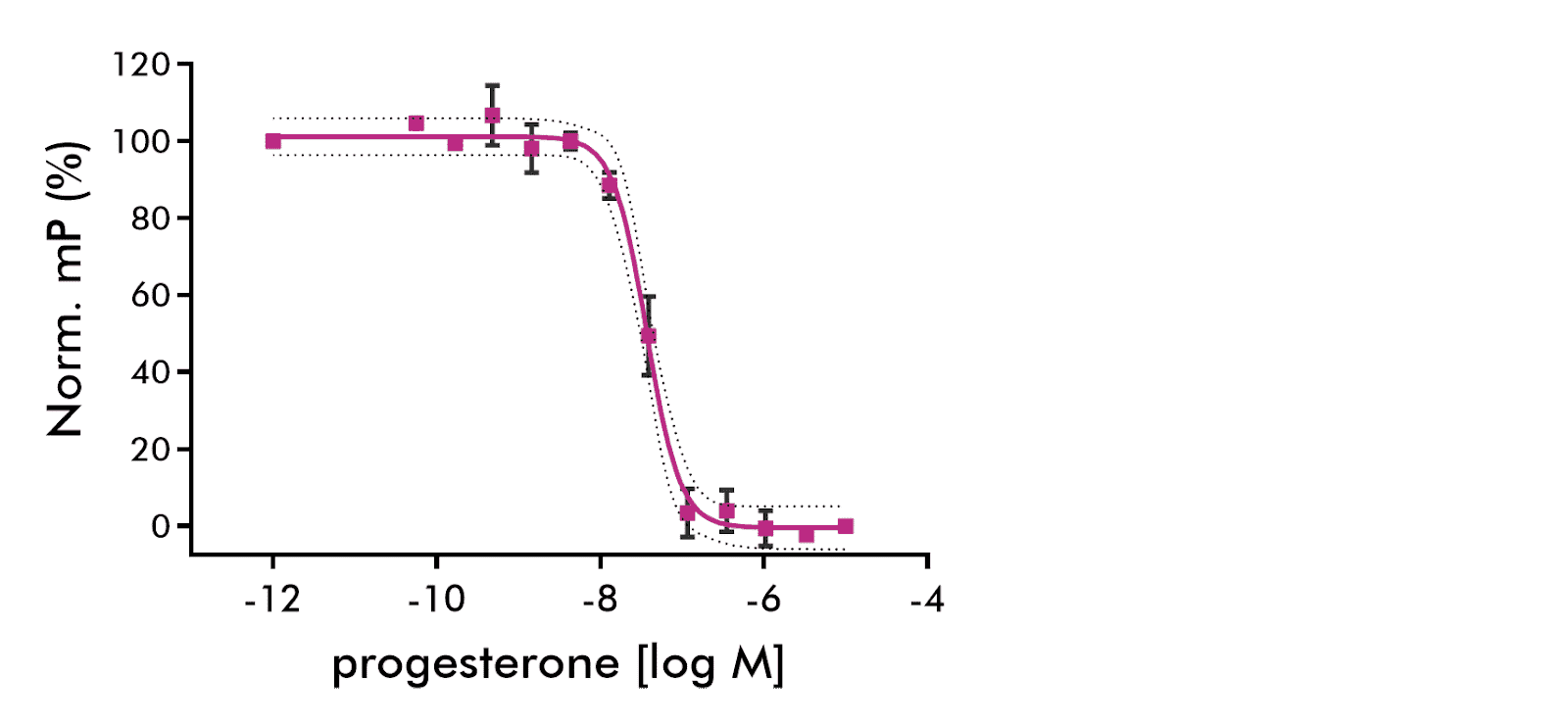
Results of three independent experiments of progesterone binding are shown. 95% CV shown as dashed lines. Sigmoid fit (solid line) parameters: IC50=36.9 nM, Hill slope=−2.13. 95% confidence intervals: EC50: 30.4 to 44.8 nM, Hillslope: ‑3.0 to ‑1.2.
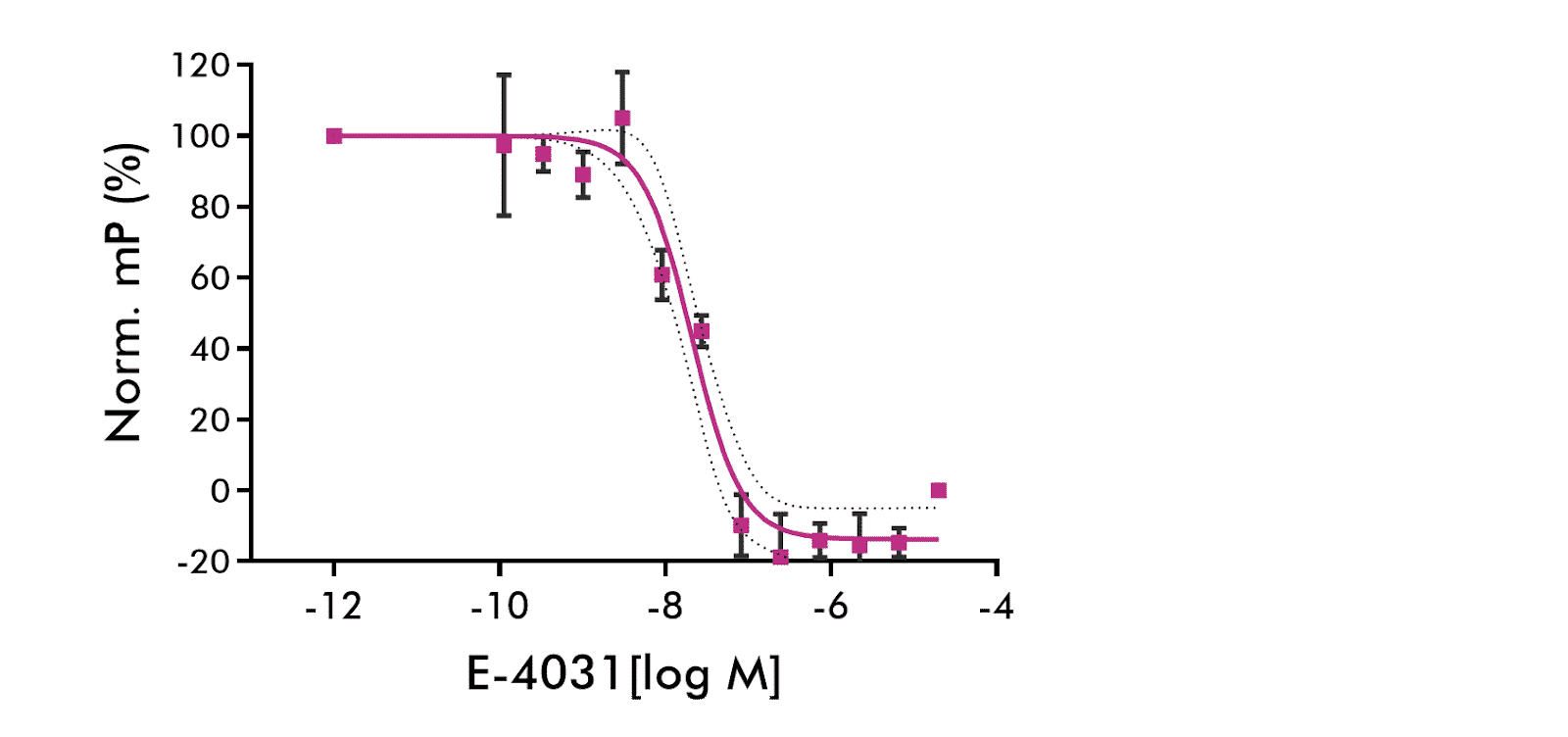
hERG Channel Fluorescence Polarization: Results of six independent experiments of E-4031 binding to hERG-containing membranes are shown. Average of 6 independent experiments. 95% CV shown as dashed lines. Competing drug: E-4031. Sigmoid fit (solid line) parameters: IC50=20.9 nM, Hill slope=−1.46. 95% confidence intervals: IC50: 14.7 to 29.7 nM, Hill slope: −2.08 to −0.84.
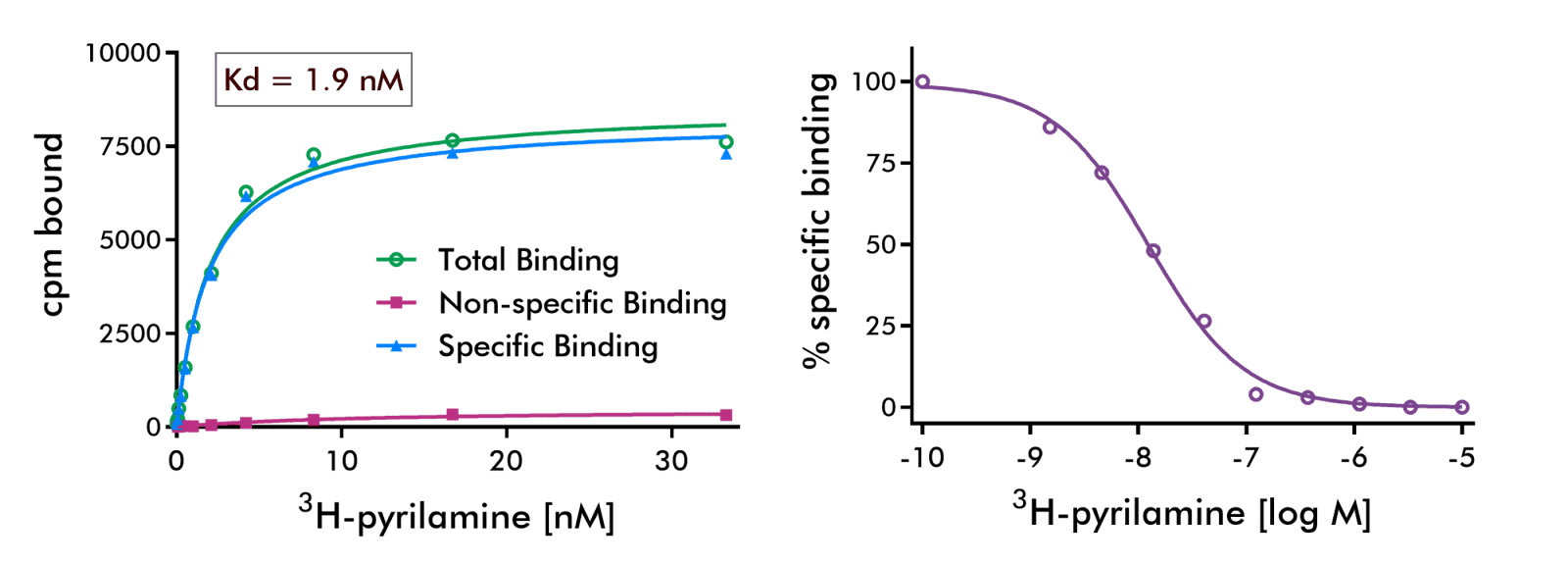
Histamine H1 Receptor Radioligand Binding Assay: Results of one experiment performed in duplicates of binding of reference compound pyrilamine to Histamine H1 receptor are shown.
Sigmoid fit parameters: EC50=1.25 nM, Hill slope=−0.99.= -0.86 Rolipram: IC50= 1.1e-06, hillslope= -0.86
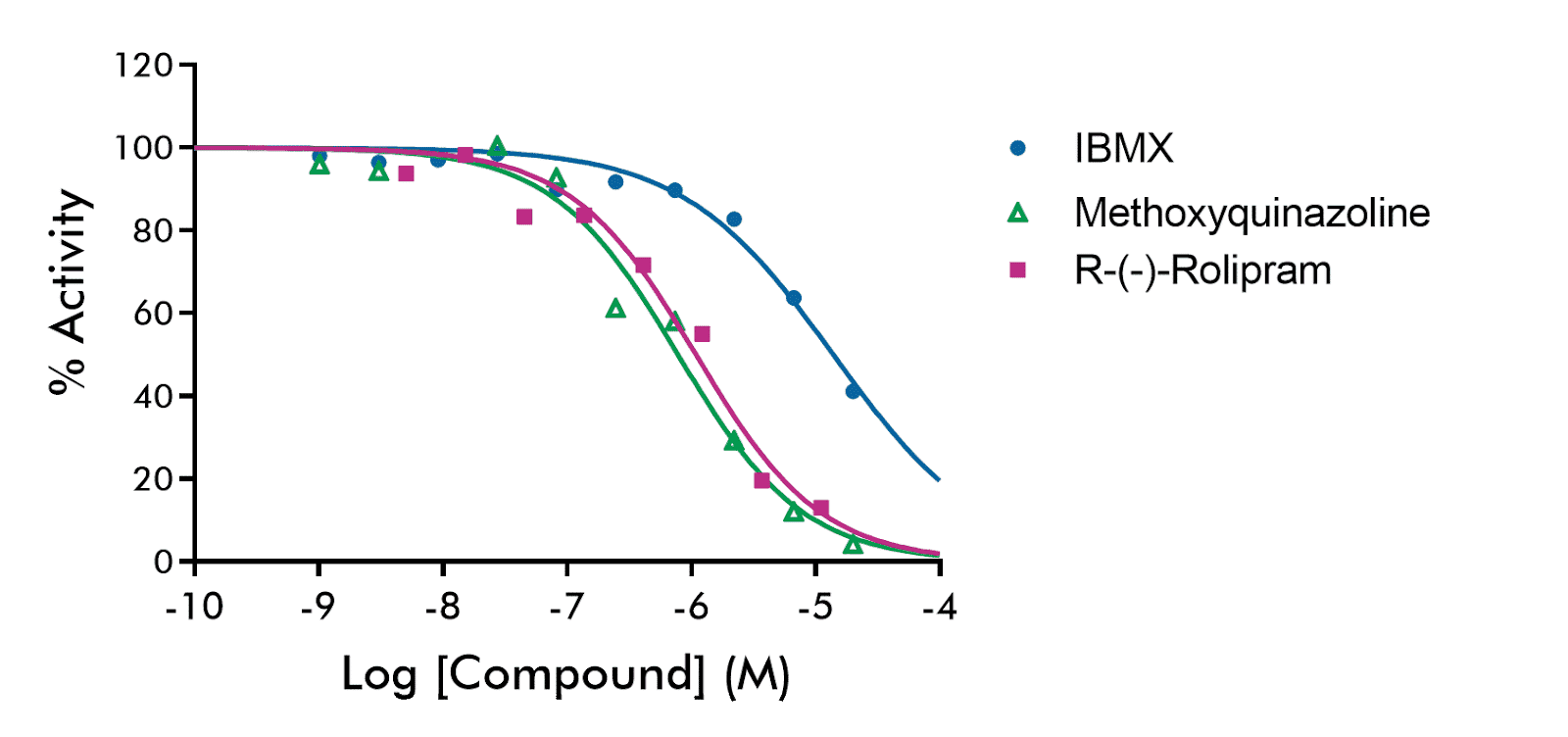
Three reference compounds IBMX, methoxyquinazoline, and Rolipram were tested against the activity of cAMP-specific cyclic phosphodiesterase 4A (PDE4A). Concentration-response curves are shown with semi-log concentrations in singlicates with the following parameters:
IBMX: IC50= 1.4e-05, hillslope= -0.72
Methoxyquinazoline: IC50= 7.82e-07, hillslope
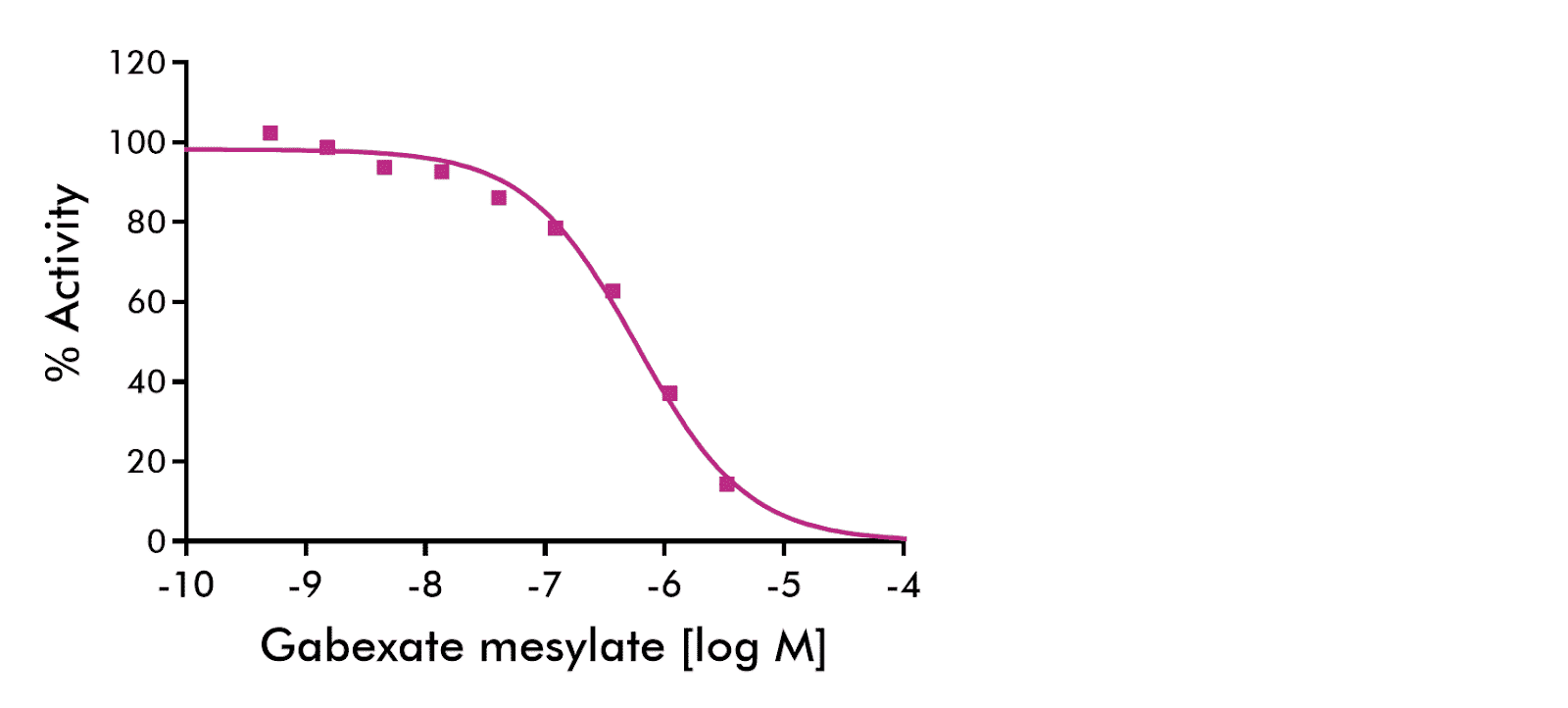
Reference inhibitor gabexate mesylate (GM) was tested against thrombin for IC50 value determination with semi-log concentrations in singlicate. The fluorescent substrate Pefafluor TH containing AMC was used as a substrate. Parameters: IC50= 5.9e-07, hillslope= -0.94
In vitro safety pharmacology—also called secondary pharmacology or off-target screening—evaluates drug candidates against receptors, enzymes, ion channels, and transporters not intended as therapeutic targets. These assays identify potential adverse drug reactions (ADRs) early in development, enabling medicinal chemistry optimization before costly in vivo studies. Standard panels screen 18–77 targets based on industry consensus (Bowes et al., 2012; Brennan et al., 2024).
Binding assays (radioligand displacement, fluorescence polarization) measure whether a compound occupies a target. Functional assays (FLIPR calcium flux, patch clamp electrophysiology, reporter gene) measure cellular response—distinguishing agonists from antagonists. InVEST panels use binding for most targets and functional assays where mechanism of action changes clinical interpretation, such as ion channels and select GPCRs.
The industry-standard 44-target panel (Bowes et al., 2012) includes 24 GPCRs (serotonin, dopamine, adrenergic, opioid, muscarinic, histamine receptors), 8 ion channels (hERG, Nav1.5, Cav1.2, KCNQ1/KCNE1, GABA(A), nAChR), 7 enzymes (AChE, COX1/2, MAO-A, PDE3A, PDE4D, LCK), 3 transporters (NET, DAT, SERT), and 2 nuclear receptors (androgen, glucocorticoid). Extended panels (59–77 targets) add kinases, additional GPCRs, and transporters.
hERG (KCNH2) encodes the cardiac potassium channel responsible for ventricular repolarization. hERG inhibition prolongs QT interval and increases risk of torsades de pointes arrhythmia—a leading cause of drug withdrawals. ICH S7B guidelines require hERG assessment. InVEST panels include hERG binding (fluorescence polarization) with functional patch clamp confirmation available for hits.
InVEST panels deliver data in 10 business days regardless of panel size (18, 44, 59, or 77 targets). This compares to 3–6 week turnaround at many providers. Monthly screening runs enable predictable scheduling; expedited timelines available for urgent programs.
Single-concentration screening (typically 10 µM, custom concentration can be requested) is cost-effective for early-stage profiling across many targets. Dose-response IC50/EC50 determination is appropriate for confirmed hits requiring quantitative potency data. InVEST offers both formats for all targets—screen broadly first, then generate curves on flagged interactions.
Most kinase inhibitors are ATP-competitive. Assays run at low ATP (Km conditions) overestimate potency because less ATP competes for the binding site. Physiological intracellular ATP is 1–5 mM. Running kinase assays at 1mM ATP provides translationally relevant IC50 values and reduces false positives that lack activity under cellular conditions. All InVEST kinase targets are profiled at 1mM ATP.
Both cover the same 44 consensus targets derived from Bowes et al. (2012). InVEST44 delivers in 10 business days from a single US facility (Malvern, PA), includes functional assays for ion channels and select GPCRs, and offers dose-response for any target. Panel selection is straightforward with no overlapping legacy nomenclature.
FDA guidance recommends screening CYP1A2, CYP2B6, CYP2C8, CYP2C9, CYP2C19, CYP2D6, and CYP3A4 for inhibition. CYP3A4 metabolizes >50% of marketed drugs. Time-dependent inhibition (TDI) screening identifies mechanism-based inhibitors with elevated DDI risk. CYP induction assessment (1A2, 2B6, 3A4) is required when inhibition or structural alerts are present. InVEST CYP covers 14 isoforms with reversible, TDI, and induction options.
The Comprehensive in vitro Proarrhythmia Assay (CiPA) paradigm evaluates hERG (IKr), Nav1.5 (INa), and Cav1.2 (ICa,L) as core channels, with KCNQ1/KCNE1 (IKs), Kir2.1 (IK1), and others for expanded assessment. InVEST includes hERG, Nav1.5, Cav1.2, and KCNQ1 via manual patch clamp. Automated patch clamp (IonFlux Mercury 16) available for higher-throughput screening.
Yes. Over 100 individual targets are available beyond standard panels. Contact with your target list to confirm availability and pricing. Custom panels can combine standard InVEST targets with additional receptors, enzymes, or kinases specific to your therapeutic area.
Gold-standard manual patch clamp for hERG, Nav1.5, Cav1.2. CiPA-aligned ion channel profiling for regulatory-grade cardiac liability data.
CYP450 inhibition and induction screening across 14 isoforms. Identify metabolism liabilities and drug-drug interaction potential early.
In vivo MTD studies to establish safe dose ranges before IND-enabling toxicology. Integrated with in vitro safety data.
Full GLP toxicology packages from our GCLP-certified Bioassay sister campus. Seamless discovery-to-IND transition.
Contact a safety pharmacology expert to discuss your in vitro safety pharmacology requirements, schedule an off-target screen, or request a custom quote for your drug safety screening panel.