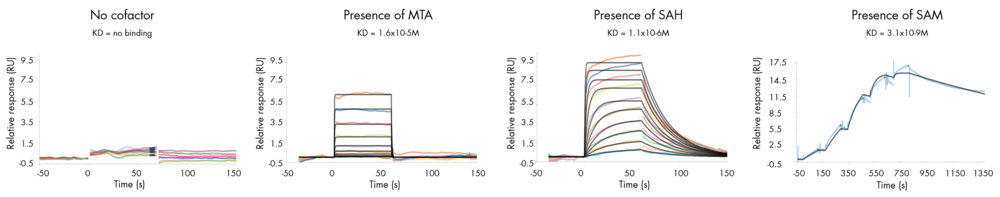Custom-tailored Epigenetic Drug Discovery
Custom-tailored epigenetic screening and mechanism of action studies can be performed with either our portfolio of assays or with an epigenetic protein or substrate provided by the customer. Customers are welcome to consult with our scientists to identify the best approach for a mechanism of action analysis based on the nature of the compound and research goals.
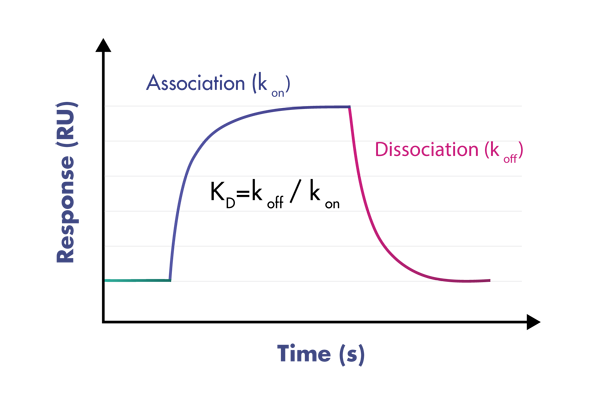
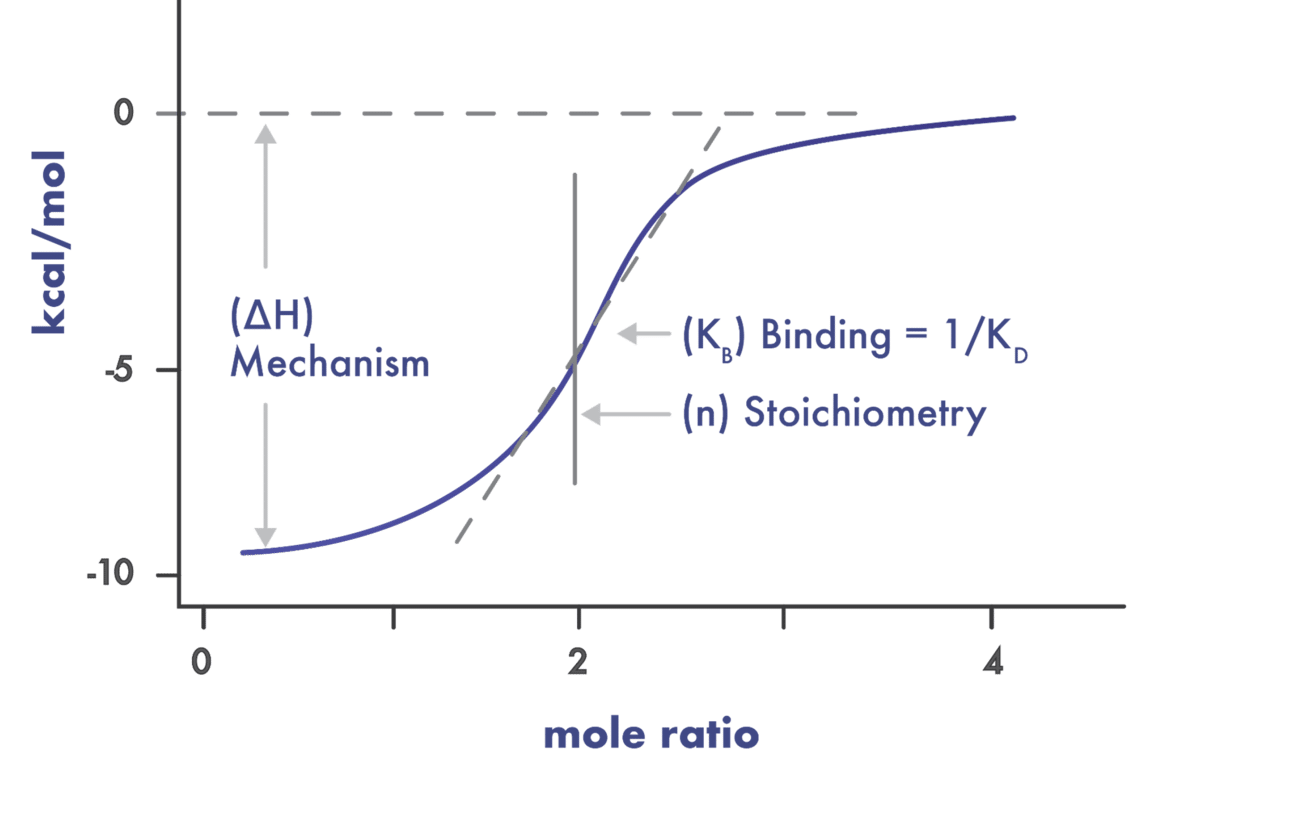
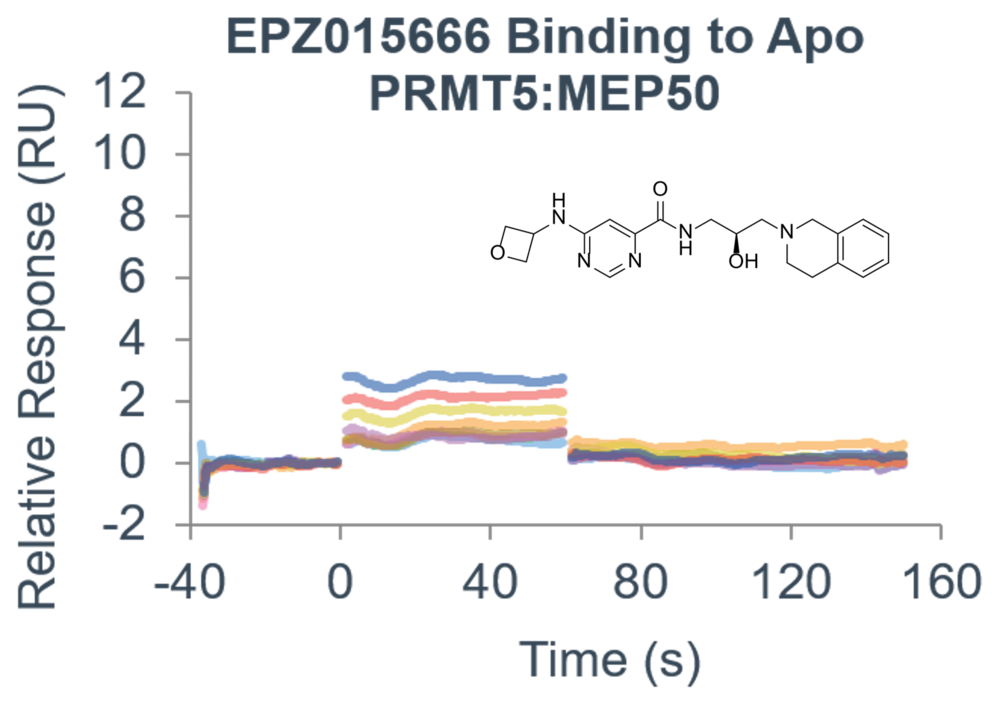
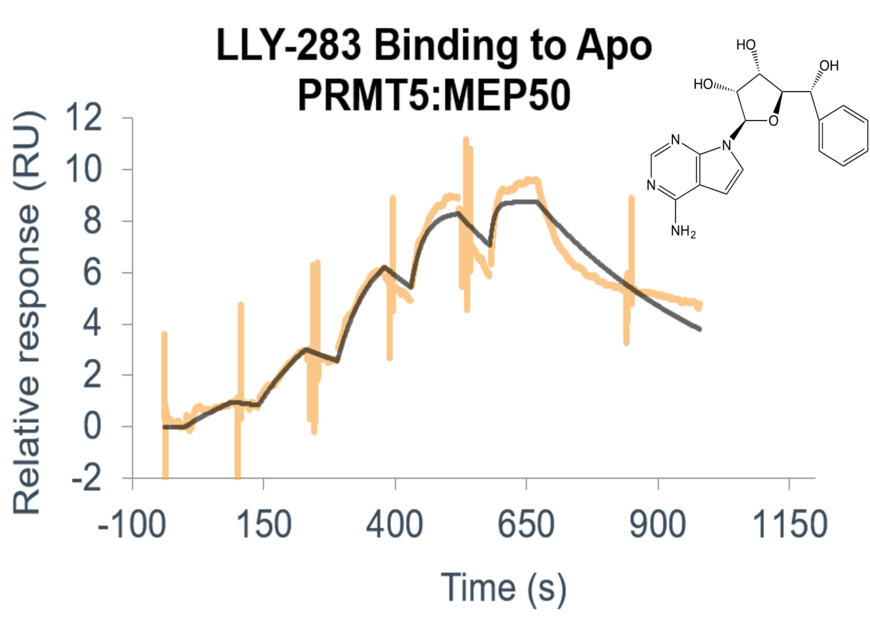
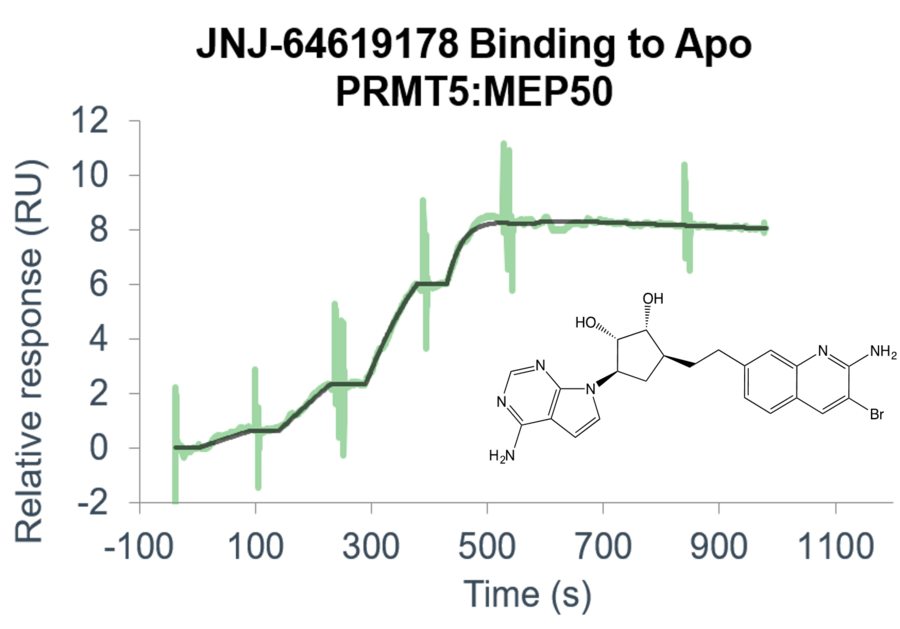
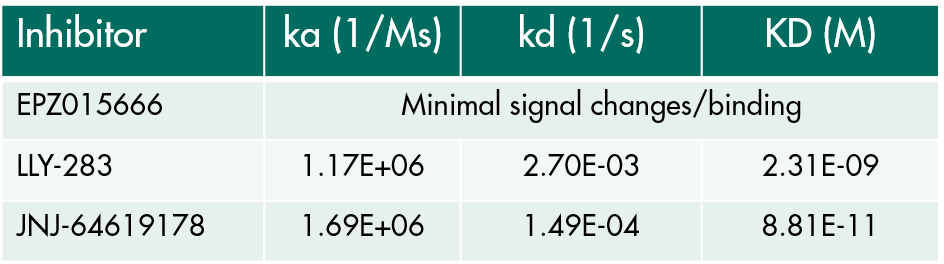
- Use a suite of biophysical assays for determination of target-compound interaction on the molecular level
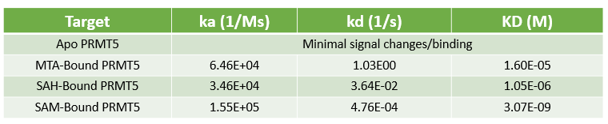
- Determine whether your compound needs the presence of a cofactor or competes with the substrate
- Custom-tailor our portfolio of epigenetic assays to suit your individual screening needs
Reaction Biology is a partner for integrated drug discovery, offering services to support every step of the drug discovery process.